ADAM User Guide¶
Introduction¶
ADAM is a library and command line tool that enables the use of Apache Spark to parallelize genomic data analysis across cluster/cloud computing environments. ADAM uses a set of schemas to describe genomic sequences, reads, variants/genotypes, and features, and can be used with data in legacy genomic file formats such as SAM/BAM/CRAM, BED/GFF3/GTF, and VCF, as well as data stored in the columnar Apache Parquet format. On a single node, ADAM provides competitive performance to optimized multi-threaded tools, while enabling scale out to clusters with more than a thousand cores. ADAM’s APIs can be used from Scala, Java, Python, R, and SQL.
The ADAM/Big Data Genomics Ecosystem¶
ADAM builds upon the open source Apache Spark, Apache Avro, and Apache Parquet projects. Additionally, ADAM can be deployed for both interactive and production workflows using a variety of platforms. A diagram of the ecosystem of tools and libraries that ADAM builds on and the tools that build upon the ADAM APIs can be found below.
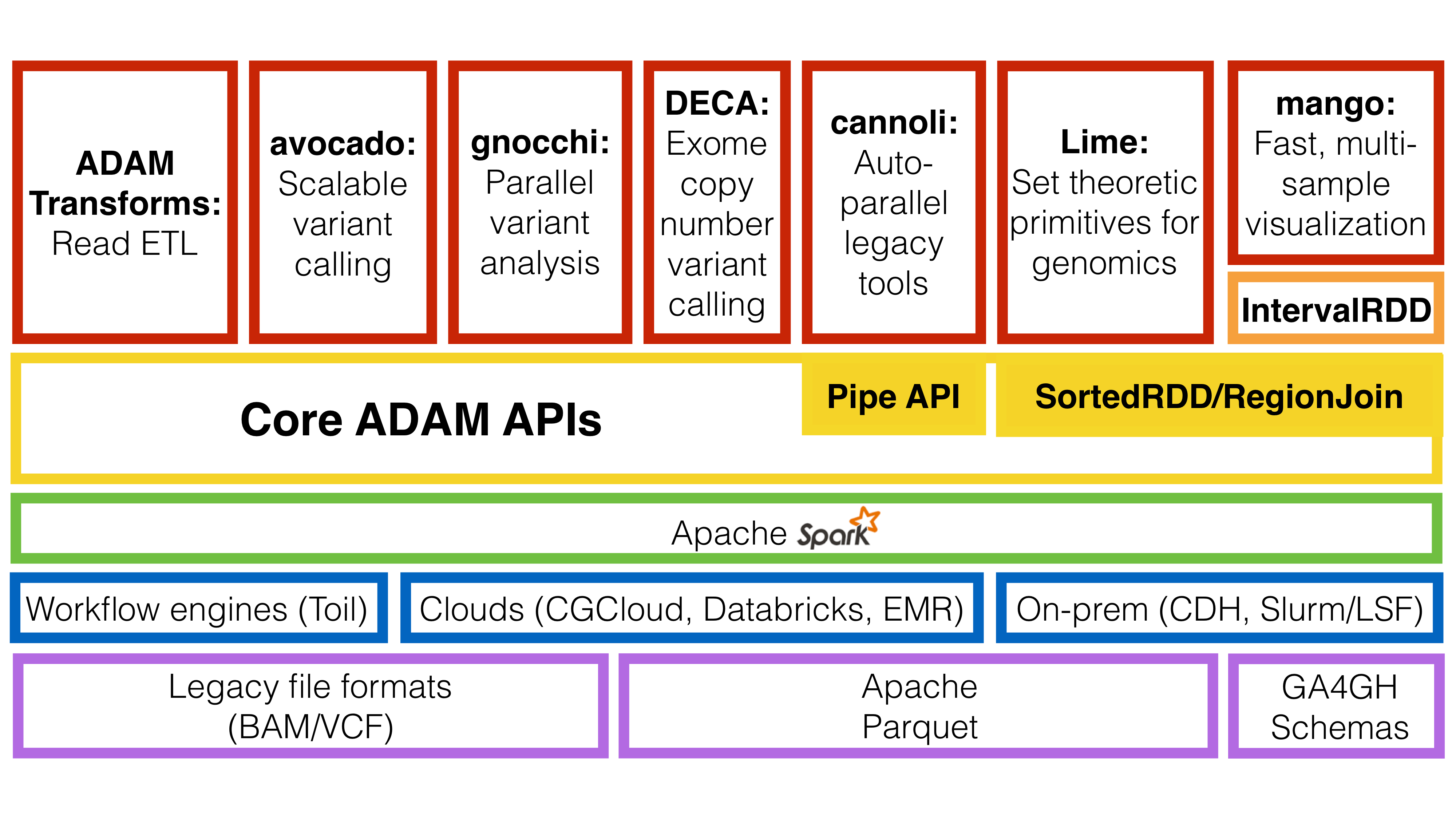
The ADAM ecosystem.
As the diagram shows, beyond the ADAM CLI, there are a number of tools built using ADAM’s core APIs:
- Avocado - Avocado is a distributed variant caller built on top of ADAM for germline and somatic calling.
- Cannoli - ADAM Pipe API wrappers for bioinformatics tools, (e.g., BWA, bowtie2, FreeBayes)
- DECA - DECA is a reimplementation of the XHMM copy number variant caller on top of ADAM.
- Gnocchi - Gnocchi provides primitives for running GWAS/eQTL tests on large genotype/phenotype datasets using ADAM.
- Lime - Lime provides a parallel implementation of genomic set theoretic primitives using the ADAM region join API.
- Mango - Mango is a library for visualizing large scale genomics data with interactive latencies.
For more, please see our awesome list of applications that extend ADAM.
Architecture
Installation
Deploying ADAM
ADAM's APIs
Building Downstream Applications
Algorithms in ADAM
References